By Ryan Prior
Every morning, Dr. David Fajgenbaum takes three life-saving pills. He wakes up his 21-month-old daughter Amelia to help feed her. He usually grabs some Greek yogurt to eat quickly before sitting down in his home office.
Then he spends most of the next 14 hours leading dozens of fellow researchers and volunteers in a systematic review of all the drugs that physicians and researchers have used so far to treat Covid-19. His team has already pored over more than 8,000 papers on how to treat coronavirus patients.
The 35-year-old associate professor at the University of Pennsylvania Perelman School of Medicine leads the school’s Center for Cytokine Storm Treatment & Laboratory. For the last few years, he has dedicated his life to studying Castleman disease, a rare condition that nearly claimed his life.
Against epic odds, he found a drug that saved his own life six years ago, by creating a collaborative method for organizing medical research that could be applicable to thousands of human diseases.
But after seeing how the same types of flares of immune-signaling cells, called cytokine storms, kill both Castleman and Covid-19 patients alike, his lab has devoted nearly all of its resources to aiding doctors fighting the pandemic.
During a cytokine storm, the body’s overactive immune response begins to attack its own cells rather than just the virus. When that inflammatory response occurs in Covid-19 patients, cytokines are often the culprit for the severe lung damage, organ failure, blood clots or pneumonia that kills them.
Having personal experience tamping down his own cytokine responses gives him a unique insight.
“I’m alive because of a repurposed drug,” he said.
Now, repurposing old drugs to fight similar symptoms caused by a novel virus has become a global imperative.

Researchers from Fajgenbaum’s lab gather in a video call to discuss Covid-19 treatment data.
A global repository for Covid-19 treatment data
Researchers working with his lab have reviewed published data on more than 150 drugs doctors around the world have to treat nearly 50,000 patients diagnosed with Covid-19. They’ve made their analysis public in a database called the Covid-19 Registry of Off-label & New Agents (or CORONA for short).
It’s a central repository of all available data in scientific journals on all the therapies used so far to curb the pandemic. This information can help doctors treat patients and tell researchers how to build clinical trials.
The team’s process resembles that of the coordination Fajgenbaum used as a medical student to discover that he could repurpose Sirolimus, an immunosuppressant drug approved for kidney transplant patients, to prevent his body from producing deadly flares of immune-signaling cells called cytokines.
The 13 members of Fajgenbaum’s lab recruited dozens of other scientific colleagues to join their coronavirus effort. And what this group is finding has ramifications for scientists globally.
Based on their database, the team published the first systematic review of Covid-19 treatments in the journal Infectious Diseases and Therapy in May.
In that first analysis of the data, the team reviewed 2,706 journal articles published on the topic between December 1, 2019, and March 27, 2020. Just 155 studies met the team’s criteria for being included in the meta-review based on standards such as the size of the cohort, the nature of the study and the end points researchers chose for concluding their inquiries.
“It’s frustrating because we all want a drug that works for everyone,” he said. But that isn’t happening because the coronavirus affects people in ways that are much more complex.
They’re sorting through oceans of data
The first key thing to consider, Fajgenbaum said, was the huge variety of Covid-19 patient experiences. It’s hard to zero in on one particular therapy because there can be such significant differences in the timing of when the drug is administered, how severely Covid-19 strikes a given individual and the stage at which the disease has progressed.
Any change in one of those variables can render an otherwise effective drug impotent. But with massive amounts of patients, the clinical data was bearing out a few noticeable themes, he said.
First, the Covid-19 patients with more severe cytokine storms were more likely to need drugs targeted toward suppressing the immune system. Those with less severe cytokine storms were likely to benefit from an immune-boosting drug.
Outside of drugs designed to boost or suppress the immune system, another major category is antiviral therapies. Various antivirals hit the “viral cascade,” Fajgenbaum said. Some work by stopping the virus from infecting cells, others by halting replication within cells. Other antivirals act in between cells and the virus.
Keeping the database is a huge undertaking, given how stunning the pace of global scientific progress and collaboration has been in the face of the disease’s human toll.
“We set the really ambitious goal of just getting this started,” Fajgenbaum said.
In the three months since the cutoff date for their first paper, the team has reviewed more than 5,000 additional papers published by scientists around the world.
One of their biggest challenges has been fitting the puzzle pieces of the different studies. With each study designed differently, one data set can’t necessarily be grafted neatly onto another. That’s especially tricky when most people diagnosed with Covid-19 eventually get better anyway. It’s hard to parse out if a particular drug was effective and saved lives.
The goal of the CORONA database isn’t to find a wonder drug per se, but to help design better clinical trials that can establish a real cause-and-effect relationship between a drug agent and an individual’s survival.
In the war against the coronavirus, Fajgenbaum hopes CORONA aims to help light the way so the heavy artillery on the front lines can better know what to shoot at Covid-19.
“It’s hard to fight a war if you’re not keeping track of what weapons are being used against the enemy,” he said.
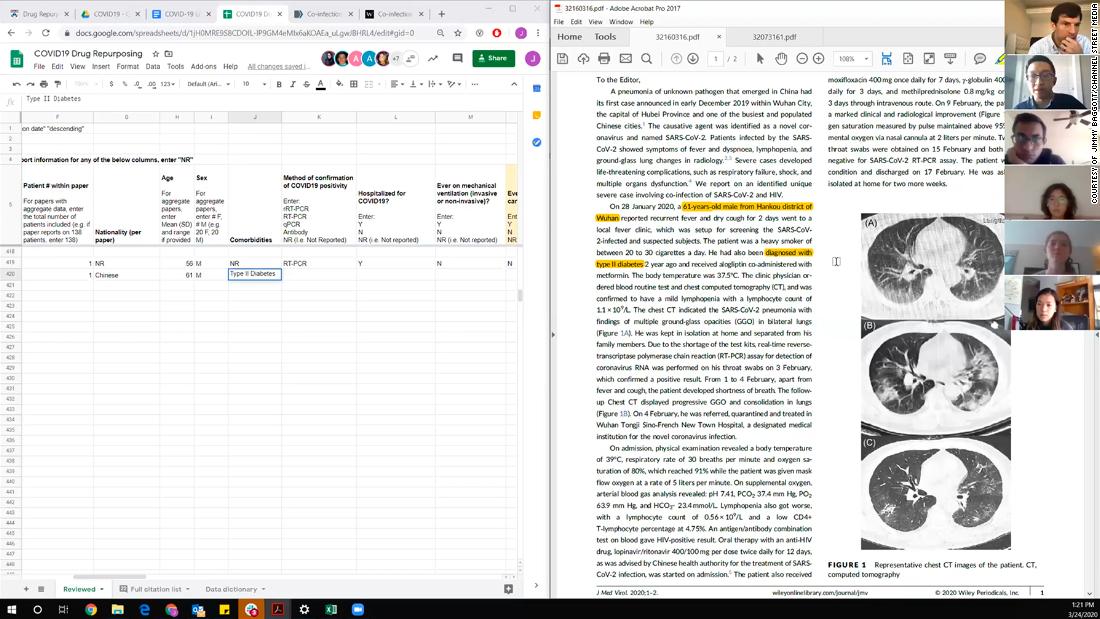
Shown here is one of the researchers’ computer screens as they review Covid-19 treatment data while on a video call. The left side shows a spreadsheet where they tabulate data from the studies. The right side shows the study they’re currently analyzing.
They’re collaborating with FDA analysts
Fajgenbaum’s CORONA database dovetails with ongoing work at the US Food and Drug Administration. For years, the agency has been developing an app called CURE ID, a platform designed to help health care providers capture novel uses of already approved drugs.
The app launched in December with two goals in mind: The first was to help advise physicians searching for new treatment ideas, prescription guidelines and emergency use advisories for drugs across hundreds of diseases. The agency’s second aim was to build a structure by which health providers in the trenches could quickly input anonymized information about their patients so that other doctors around the world could quickly see whether they had been successful using an off-label drug.
The app was ready just in time for the pandemic, and Fajgenbaum gave the keynote speech at its launch.
“It’s really been a terrific collaboration,” said a health policy analyst with the FDA. “His life follows very much the model we hope to use.”
Now that he and his team are working on the coronavirus, the urgency of their partnership has strengthened.
“Nobody wants to go to a database with no data in it,” the analyst said. “Rather than reinventing the wheel, he was kind enough to provide all his data.”
And while the CORONA database project is primarily intended to aid researchers, it’s tapping into major currents in health economics that explain weak points in the way the public and private sector develop therapies together.
“Covid-19 illustrates a market failure in how we build vaccines,” said Amitabh Chandra, a health economist with joint appointments as a professor at the Harvard Kennedy School and Harvard Business School. “We haven’t given firms the correct incentives to make vaccines before a pandemic. Vaccines are very hard to test before the pandemic hits.”
There aren’t old vaccines sitting on a shelf waiting to be dusted off to save the world from the coronavirus. But there are hundreds of FDA-approved drugs at your local pharmacy that can save lives immediately.
When teaching classes, Chandra uses a 2017 New York Times story profiling Fajgenbaum to illustrate the value of drug repurposing and motivate his students to think boldly about how to create economic incentives to cure diseases, particularly when a “invisible medicine” might be right under your nose.
“There’s no substitute for a good story to get people motivated,” he said.
Many drugs are beginning to stand out.
The combination of antivirals lopinavir and ritonavir is the Covid-19 treatment protocol with the most number of studies published so far. As of mid-June, the team had looked at papers on that drug pairing involving more than 4,500 patients.
Next, corticosteroids have shown particular promise, making appearances in studies with another 4,000 patients. At the cellular level, antivirals work for a variety of reasons, each with its own specialty in attacking the virus at different points in its life cycle. Corticosteroids are different, however.
“Steroids tend to act the same, with replicating cortisol,” Fajgenbaum said.
He feels particularly elated about a recent United Kingdom-based study on the steroid dexamethasone. The study garnered headlines for its result showing that a low-dose 10-day regimen of the drug could reduce the risk of death by a third among hospitalized patients requiring ventilation.
In their spreadsheets, the numbers around dexamethasone were like a beacon.
“We built CORONA to help uncover something like dexamethasone,” he said. “It’s a cheap repurposed drug that’s been around for 60 years. This is what it’s all about.”
Studies need rigor
Because Covid-19 is so new, many of the studies are observational or anecdotal. These types of studies obviously matter as scientists are building a foundation of knowledge.
But the best insights come from running double-blind placebo-controlled studies. One shortfall is that many of the published studies just don’t have the level of rigor to inform larger-scale scientific decision-making.
“There are a lot of biases in these observational studies,” Fajgenbaum said.
One drug, the anti-malarial drug hydroxychloroquine, has famously received a lot of boosterism from US President Donald Trump. But in the published studies available for Fajgenbaum’s team to review, the drug hasn’t outperformed others.
Two French studies on hydroxychloroquine drew red flags for the University of Pennsylvania-based team because of the clinical end point the researchers chose: the time when the coronavirus cleared the body. It can be problematic to base an argument for a drug’s success only on that particular metric, because it leaves out crucial details from a person’s longer-term experience following infection.
“‘Virally cured’ is a challenging term,” Fajgenbaum said. “We don’t know if they’re discharged how they fared after leaving the hospital.”
On top of that, the reviewers were skeptical because the virus took a long time to leave the patients’ bodies, which they refer to as “a high time to viral clearance.”
That indicator that could suggest the drug was slow to take effect, or that other factors, including the patient’s own immune system, played a larger role in expelling the pathogen.
Know how to sort through the data
With dozens of people working full time to sort through thousands of studies, it’s obviously impossible for a single frontline health provider to keep abreast of all there is to know about Covid-19 while also treating patients at the same time.
It’s even harder for the average person following the story in the news, especially if you’re not equipped with a graduate degree in statistical analysis.
“Covid threw the world in flux,” said Sheila Pierson, associate director for clinical research at the CSTL. A biostatistician originally hired to study Castleman disease, she’s accepted the new mission along with her colleagues.
“There’s a lot of great science being done,” she explained. With that pace of innovation, it’s incredibly difficult for the average person to stay up to date, so the CORONA database helps everyone with a little extra scientific literacy amid headlines about new treatments that induce a form of intellectual whiplash.
“You should rely on multiple news sources,” Pierson said, in order to sort through what may appear to be conflated messages about whether a certain drug works or not for a certain group of people.
“It’s difficult when you’re only looking at one person’s view of a drug,” she said. “Look for a different write-up and a different view.”
He’s repeating the same methods that saved his life
As of June 27, Fajgenbaum has lived free of Castleman’s cytokine storms for 77.72 months. His last Castleman relapse ended on January 5, 2014. He’s a living experiment, and in his personal accounting he won’t round up to the next full month. Each new day is a precious moment with a daughter he feared he’d never meet.
The doctor and researcher remains immune compromised and won’t take risks with the coronavirus.
He hasn’t set foot in a building other than his home since March 13. And his life still relies on siltuximab and chemotherapy infusions administered monthly through a chest port.
“I’m reminded every time I touch the port in my chest of the cytokine storms I had,” he said. “I want so badly to solve (Covid-19) the way I did with Castleman. I have the same sense of urgency.”
Castleman disease nearly killed Fajgenbaum five times in his 20s while he was working his way through University of Pennsylvania’s Perelman School of Medicine and then earning an MBA at the University of Pennsylvania’s Wharton School.
Each time, the deadly disease triggered cytokine storms that led to multiple organ failure.
But the young man created a global organization to rally doctors, scientists and patients toward finding a cure. With intense study and brilliant partners, he zeroed in on an already available immunosupressant that could be repurposed to save his life.
Last year he published his memoir, “Chasing My Cure,” detailing a journey in which at one point a priest was brought to his hospital room to give his last rites.
Fajgenbaum’s story reads likes the teaser for a hit Netflix series. But if it were a show, all of that is really just season one. Because, spoiler alert — then a global pandemic hit.
A year ago you might have thought what the writers threw at him in a second season might be a bit unrealistic. But this project is the obvious next step.
“I see myself bringing our experiences with Castleman now over to the global fight against corona,” he said.
https://www.cnn.com/2020/06/27/health/coronavirus-treatment-fajgenbaum-drug-review-scn-wellness/index.html