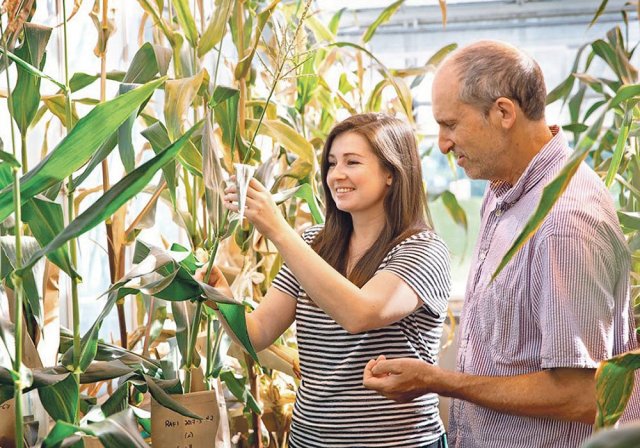
Professor David Stern examines test corn plants at Boyce Thompson Institute with Coralie Salesse-Smith. They are looking for varieties that will be better able to cope with cold weather. | Boyce Thompson Institute photo
Corn is the third largest grain crop in Canada and the number one crop in Ontario in terms of production. Nationally, yields vary depending on weather, and 2019 was a particularly challenging year.
Heavy rain and colder than normal temperatures made planting conditions difficult in Eastern Canada. Very dry conditions persisted into the growing season in parts of Western Canada, while unseasonal rain and early snow on the Prairies slowed harvest for many farmers.
Corn is one of the world’s most important crops, not only for food but for animal feed and biofuel. However, as a grass of tropical origin, it is particularly sensitive to cold weather. That trait can be problematic, especially when the growing season is only four or five months.
At Cornell University’s Boyce Thompson Institute in New York, scientists have developed a chill-tolerant corn variety that recovers much more quickly after a cold snap. It could broaden the latitudes in which corn can be grown and help farmers increase yields, especially in the face of wildly fluctuating weather patterns due to climate change.
“While the research is too early-stage to know for sure, the chilling-tolerant corn has promise to help crops cope with early cold snaps, as well as recover more quickly from drought,” said David Stern, president of the Boyce Thompson Institute and adjunct professor of plant biology in Cornell University’s College of Agriculture and Life Sciences.
“In terms of cold snaps, the combination of cold weather and strong spring sunlight can cause corn leaves to bleach, weakening or even killing the plants. The chilling-tolerant corn is less prone to bleaching, allowing it to recover more quickly. As a result, farmers could potentially plant earlier and harvest earlier, avoiding the frequent drought-prone conditions of late summer that occur just when ears are filling with grain. The potential benefits are a more flexible spring planting schedule, and an earlier harvest.”
Research in 2018 showed that increasing levels of an enzyme called rubisco led to bigger and faster-maturing plants. Rubisco is needed by plants to turn atmospheric carbon dioxide into sugar but, in cold weather, its levels in corn leaves decrease dramatically.
“Plants are very good at sensing temperature and seem to deliberately reduce the amount of rubisco when it’s cold,” said Stern.
“In doing so, plants can save the energy it takes to make rubisco and other proteins, as well as generally slow down metabolism, just like many living organisms do in the cold (think of hibernation).”
Rubisco is made up of proteins — eight large subunits and eight small ones with help from a third protein called RAF1. Because rubisco is a protein, it is made up of amino acids, which are rich in nitrogen, making rubisco a nitrogen reservoir. However, Rubisco is actually an inefficient enzyme so boosting its function will boost plant growth.
“We introduced transgenes that increase the abundance of (the) three proteins: the large subunit of rubisco, the small subunit and RAF1,” said Stern.
“In the modified corn, all three proteins are more abundant, thus increasing the level of Rubisco.”
The scientists grew plants for three weeks at 25 C, lowered the temperature to 14 C for two weeks and then returned it to 25 C, which is a considerable temperature swing. The intent was to test resistance and monitor the factors that can help plants withstand stress.
“The corn with more rubisco performed better than regular corn before, during and after chilling,” said Coralie Salesse-Smith, a PhD candidate in Stern’s lab during the study and the research paper’s first author.
“We were able to reduce the severity of chilling stress and allow for a more rapid recovery.”
Compared to traditional corn, the genetically modified corn had higher photosynthesis rates throughout the experiment and recovered more quickly from the chilling stress with less damage to the molecules responsible for photosynthesis. As a result, the plant grew taller and developed mature ears more quickly following a cold spell.
“What scientists are trying to build into plants is resilience — the ability to withstand a variety of shocks whose frequency and magnitude cannot be reliably predicted,” said Stern.
“In the laboratory, we can only test a small number of simulated climactic conditions. What happens when cold is combined with drought? Flooding with a heat wave? The real world of agriculture is far more complicated than a lab study. So, putting this kind of corn in the field is the best way to test its resilience.”
He said that the technology to bolster corn into a chill-resistant variety is being tested for the first time this growing season by a large seed company. They will know later this year or early next year if the plant is responding the way they expect and if it works as well in their elite field varieties as it does in the laboratory.
“If it does, the traits would move into their standard breeding pipeline and farmers would see the seed in six to 10 years. That is the reality of breeding for new corn traits,” said Stern.
Meanwhile, several research projects continue.
“One is to improve the activity, or speed, of the rubisco enzyme,” he said.
“In corn, only about 70 percent of the enzyme is working at any given time. We would like to increase that to 80 percent or 90 percent. Another path is to combine the rubisco trait with other genes that can increase photosynthesis. Rubisco is only one player in a complicated metabolic network, and we need to target other steps in the pathway as well. A third direction is to combine this higher photosynthesis-chilling tolerance trait with genes that improve tolerance to insects, drought or heat. This moves towards the ‘optimal’ resilient and climate-ready plant that would be a long-term goal.”
It is possible this approach could be used on other crops such as sugar cane and sorghum. The research paper was published online in the Plant Biotechnology Journal.
https://www.producer.com/2020/04/new-corn-variety-copes-better-with-cold/