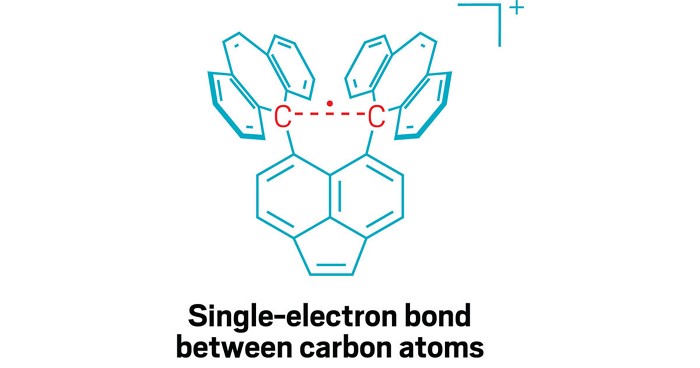
For a little more than a century, chemists have believed that strong atomic links called covalent bonds are formed when atoms share one or more electron pairs. Now, researchers have made the first observations of single-electron covalent bonds between two carbon atoms.
This unusual bonding behaviour has been seen between a few other atoms, but scientists are particularly excited to see it in carbon, the basic building block of life on Earth and the key component of industrial chemicals including drugs, plastics, sugars and proteins. The discovery was published1 in Nature on 25 September.
“The covalent bond is one of the most important concepts in chemistry, and discovery of new types of chemical bonds holds great promise for expanding vast areas of chemical space,” says University of Tokyo chemist Takuya Shimajiri, who was part of the carbon bonding research team.
Most chemical bonds in molecules are made up of a single pair of electrons, shared between atoms. These are called covalent single bonds. In particularly strong bonds, atoms might share two electron pairs in a double bond, or three pairs — a triple bond. But chemists know that atoms interact in many other ways, and by studying more unusual bond types at the boundaries of the possible, they hope to better understand what a chemical bond is in the first place.
Pauling’s proposal
The concept of single-electron covalent bonds dates to 1931, when chemist Linus Pauling proposed them. But at the time, chemists didn’t have the tools to observe such bonds, says Marc-Etienne Moret, a chemist at Utrecht University in the Netherlands. Even with modern analytical techniques, these bonds are challenging to observe. “The situation in which only one electron makes a bond is very unstable,” says Moret. “This means the bond will break easily and have a strong tendency to either release or capture an electron to restore an even number of electrons.”
In 1998, scientists observed2 a single-electron bond between two phosphorus atoms; Moret was part of a group that created3 one between copper and boron in 2013. Chemists have theorized that these unusual bonds might occur between carbon atoms in short-lived intermediate structures that appear during chemical reactions. But to observe these fickle bonds, chemists have to stabilize a compound that contains them. A stable compound that contains a one-electron C–C bond had eluded chemists.
Shimajiri says the key to observing the single-electron carbon bond was carefully designing a molecule that would stabilize it. The research team, which included Hokkaido University chemist Yusuke Ishigaki, created a molecule that provides a stable ‘shell’ of fused carbon rings that helps hold together the carbon–carbon bond in its centre. That central bond is stretched out to a relatively long length for a C–C bond, which makes it susceptible to losing one electron in an oxidation reaction, creating the elusive single-electron bond.
Stable bond
To capture this compound in a stable, observable form, they crystallized it. When the oxidation is performed in the presence of iodine, the reaction yields a purple salt, with the stable shell of the molecule holding together the single-electron C–C bond inside. They then used various analytical techniques to characterize the molecule and the bond. Shimajiri says the compound is extremely stable under ambient conditions.
“In several chemical reactions, the involvement of one-electron bonds has been proposed, but so far, they have remained hypothetical,” says Shimajiri. Creating stable compounds containing these bonds could help researchers to better understand what happens during these reactions.
Guy Bertrand, a chemist at the University of California, Santa Barbara, was part of the team that created the phosphorus single-electron bond. He says it’s significant to see it in carbon. “Anytime you do something with carbon, the impact is greater than with any other element,” he says. Carbon is the stuff of organic chemistry. But he says it’s not so easy to say whether this work will have any applications. “This is a curiosity,” he says. “But it will be in the textbooks.”
Shimajiri hopes that the description of the single-electron carbon bond will help chemists to better understand the basic nature of chemical bonds. “We aim to clarify what a covalent bond is — specifically, at what point does a bond qualify as covalent, and at what point does it not?”
doi: https://doi.org/10.1038/d41586-024-03138-2

















