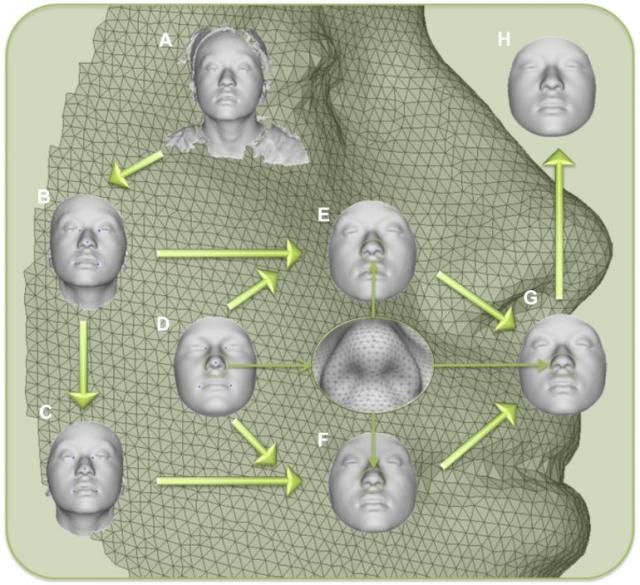
Workflow for 3D face scan processing, including the A) original surface, B) trimmed to exclude non-face parts, C) reflected to make mirror image, D) anthropometric mask of quasi-landmarks, E) remapped, F) reflected remapped, G) symmetrized and H) reconstructed.
By Philip Ross
Could a single hair be used to make an accurate 3D model of a criminal suspect’s face? Researchers from the U.S. and Belgium have developed a computer program that renders a crude genetic “mugshot” from a small sample of DNA.
Forensics can already predict eye and hair color relatively easily. Io9 notes that criminal investigators can even use maggots to extract a victim’s DNA from their unidentifiable body or find hidden faces by zooming into hi-res photos of eyes. But the face is a complex structure that’s more difficult to map from just one DNA sample.
According to New Scientist, researchers used a stereoscopic camera to make 3D images of roughly 600 volunteers with mixed European and West African ancestry. They identified more than 7,000 distinct points on the face to see how sex and racial ancestry affect the position of these points. The variations were used to develop a statistical model that reconstructs the overall shape of a person’s face.
The team also isolated 24 genetic variants, called single nucleotide polymorphisms, which play a role in shaping a face, such as those that shape the head during embryonic development. Lastly, researchers had volunteers rate the 600 faces on perceived ethnicity as well as on a scale of masculinity and femininity.
The new study, published in the journal PLOS Genetics, says this process could allow investigators to make computer-generated mugshots from genetic material left at a crime scene.
“We show that facial variation with regard to sex, ancestry, and genes can be systematically studied with our methods, allowing us to lay the foundation for predictive modeling of faces,” the authors note. “Such predictive modeling could be forensically useful; for example, DNA left at crime scenes could be tested and faces predicted in order to help to narrow the pool of potential suspects. Further, our methods could be used to predict the facial features of descendants, deceased ancestors, and even extinct human species. In addition, these methods could prove to be useful diagnostic tools.”
Any 3D renderings created using the new technology wouldn’t be used in a court of law – any person identified via the DNA mugshots would still have his DNA compared to the crime scene sample – but it could at least narrow the search for a suspected criminal. And there are still a few kinks to work out in the process before the technology is ever used in the field.
“I believe that in five to 10 years’ time, we will be able to computationally predict a face,” Peter Claes of the Catholic University of Leuven in Belgium told New Scientist.
Thanks to Da Brayn for bringing this to the attention of the It’s Interesting community.

